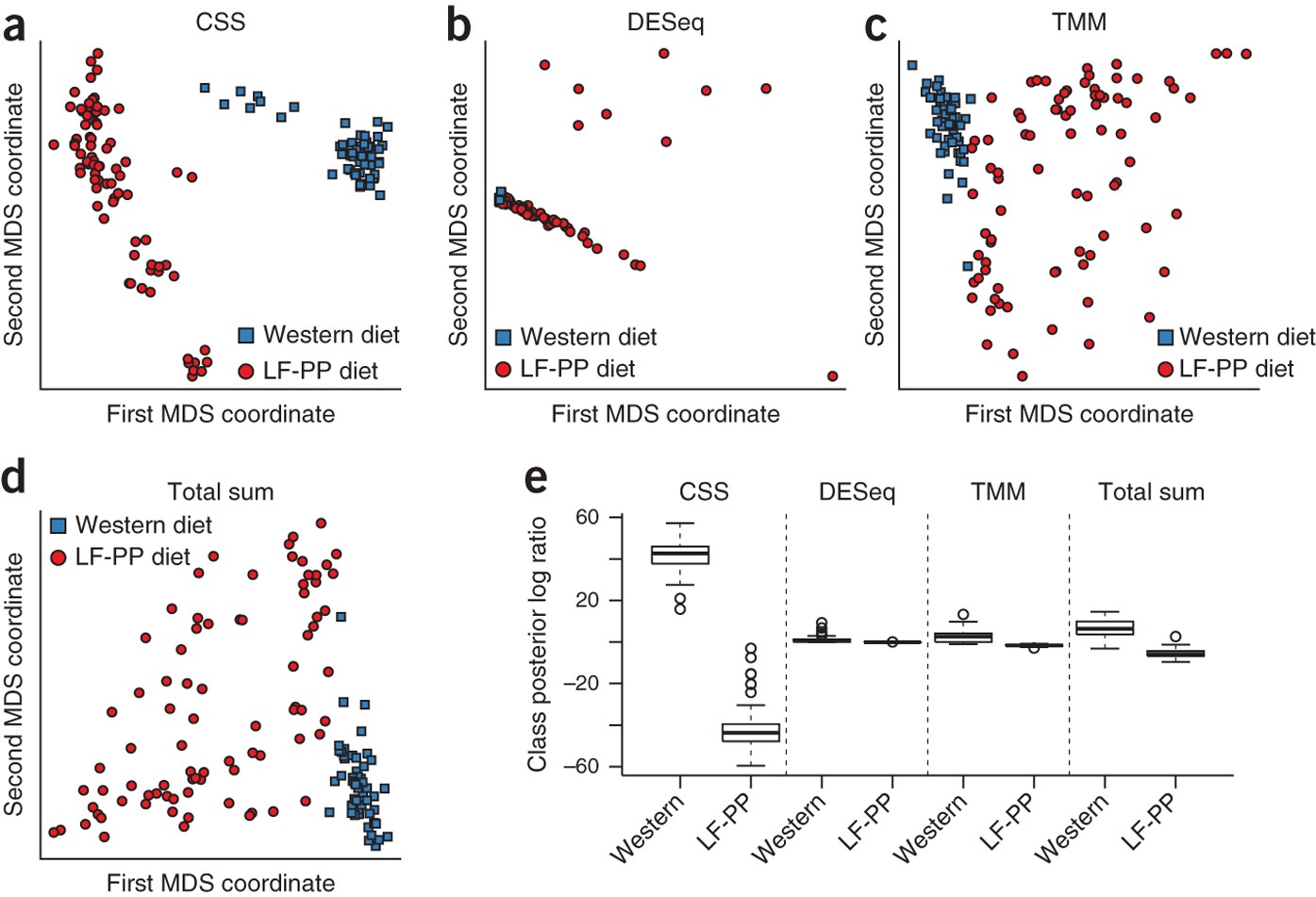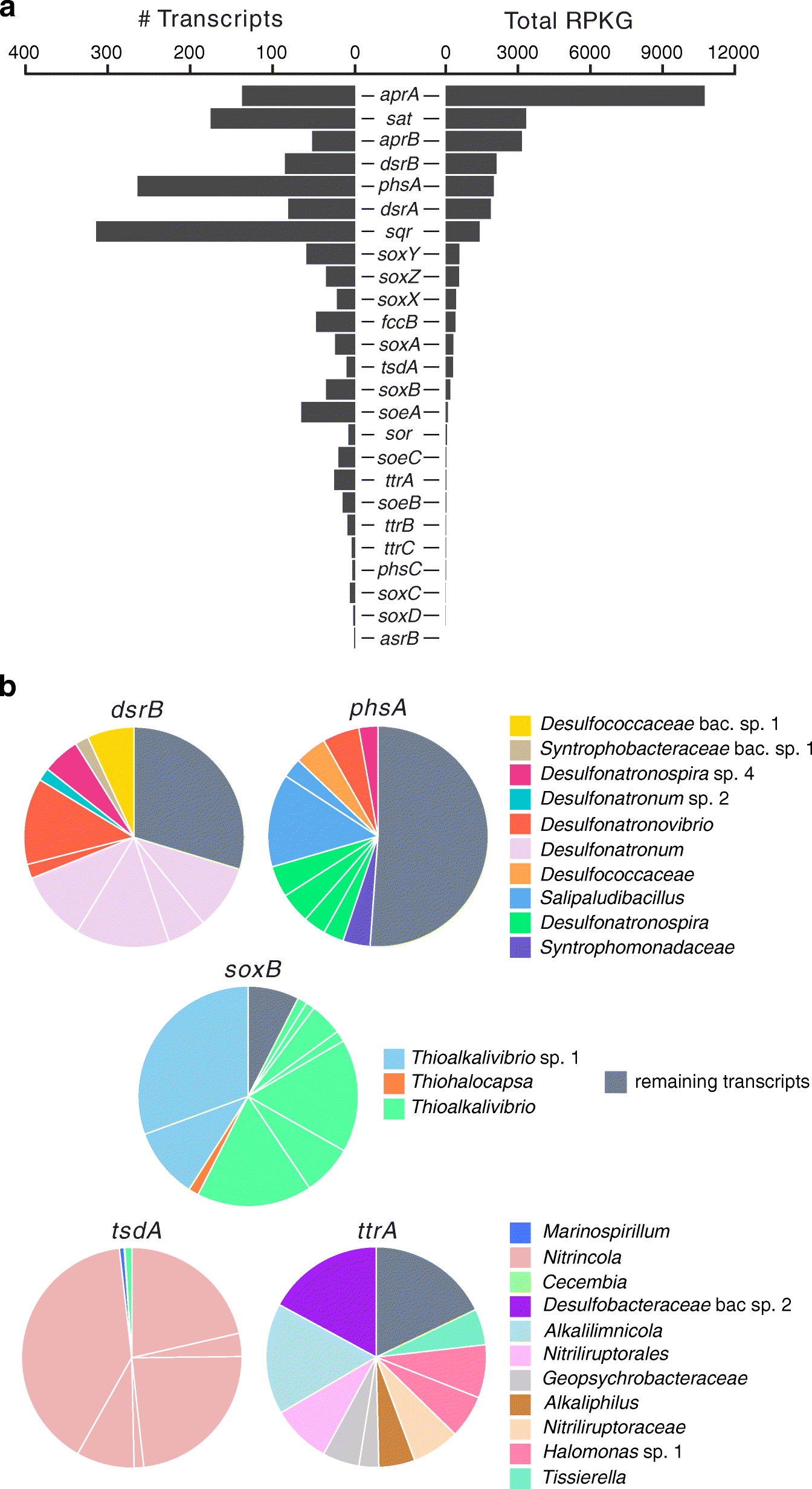
Metagenomes and metatranscriptomes shed new light on the microbial-mediated sulfur cycle in a Siberian soda lake | BMC Biology | Full Text

Virus-inclusive single-cell RNA sequencing reveals the molecular signature of progression to severe dengue | PNAS
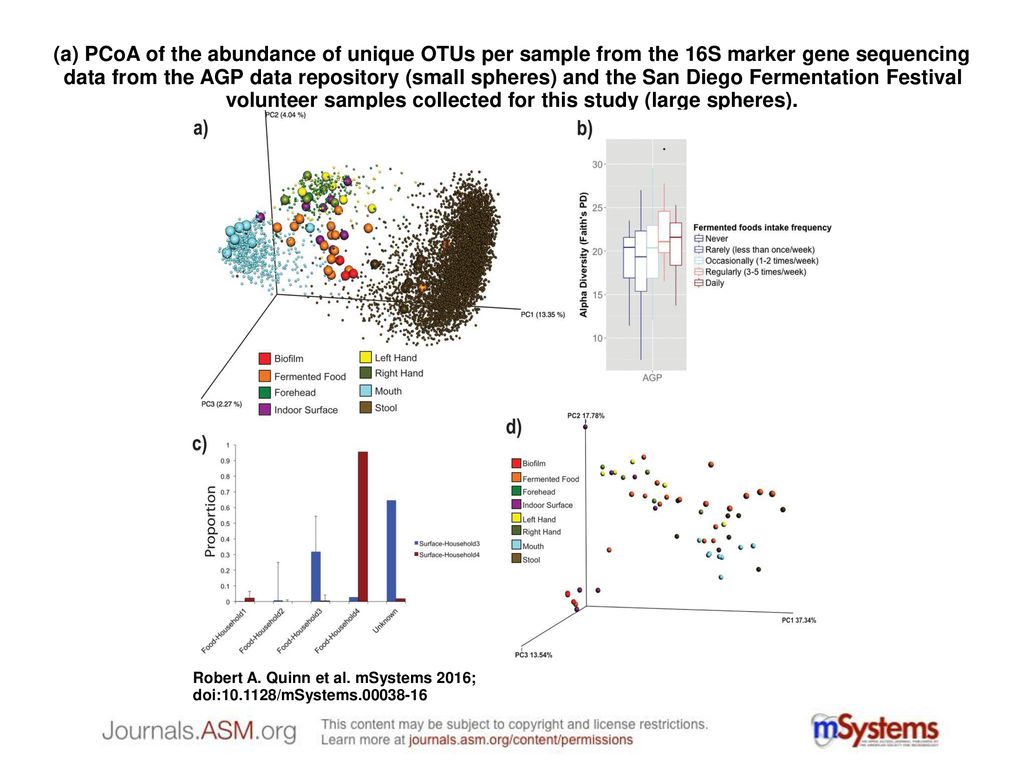
a) PCoA of the abundance of unique OTUs per sample from the 16S marker gene sequencing data from the AGP data repository (small spheres) and the San Diego. - ppt download

A neural network‐based framework to understand the type 2 diabetes‐related alteration of the human gut microbiome - Guo - 2022 - iMeta - Wiley Online Library
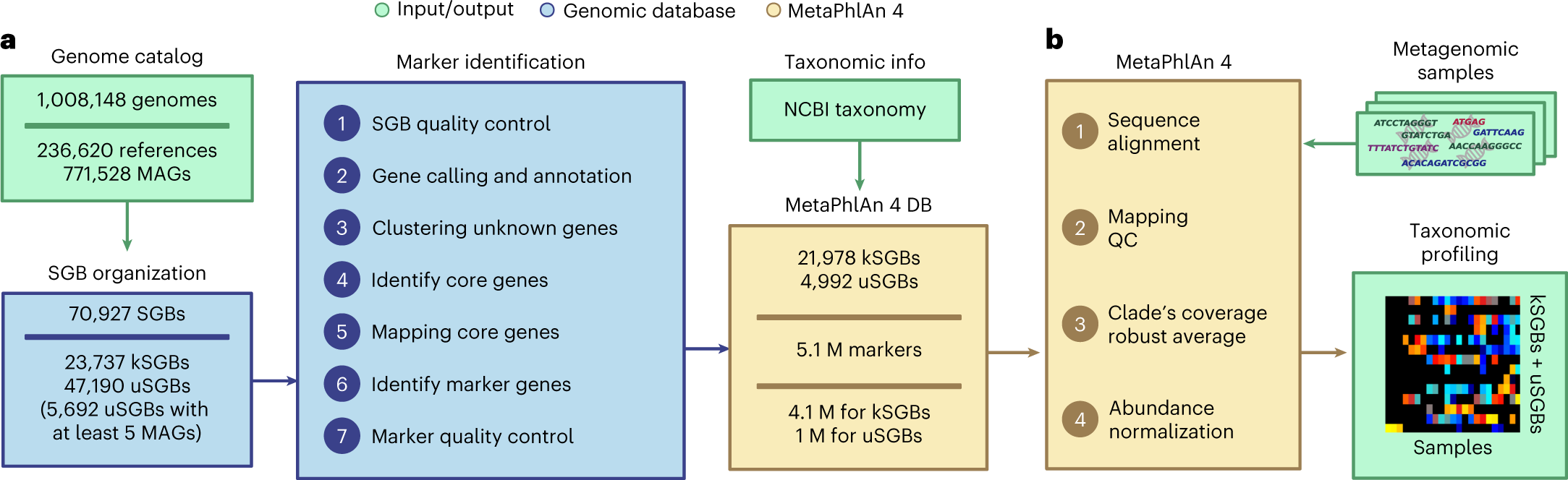
Extending and improving metagenomic taxonomic profiling with uncharacterized species using MetaPhlAn 4 | Nature Biotechnology
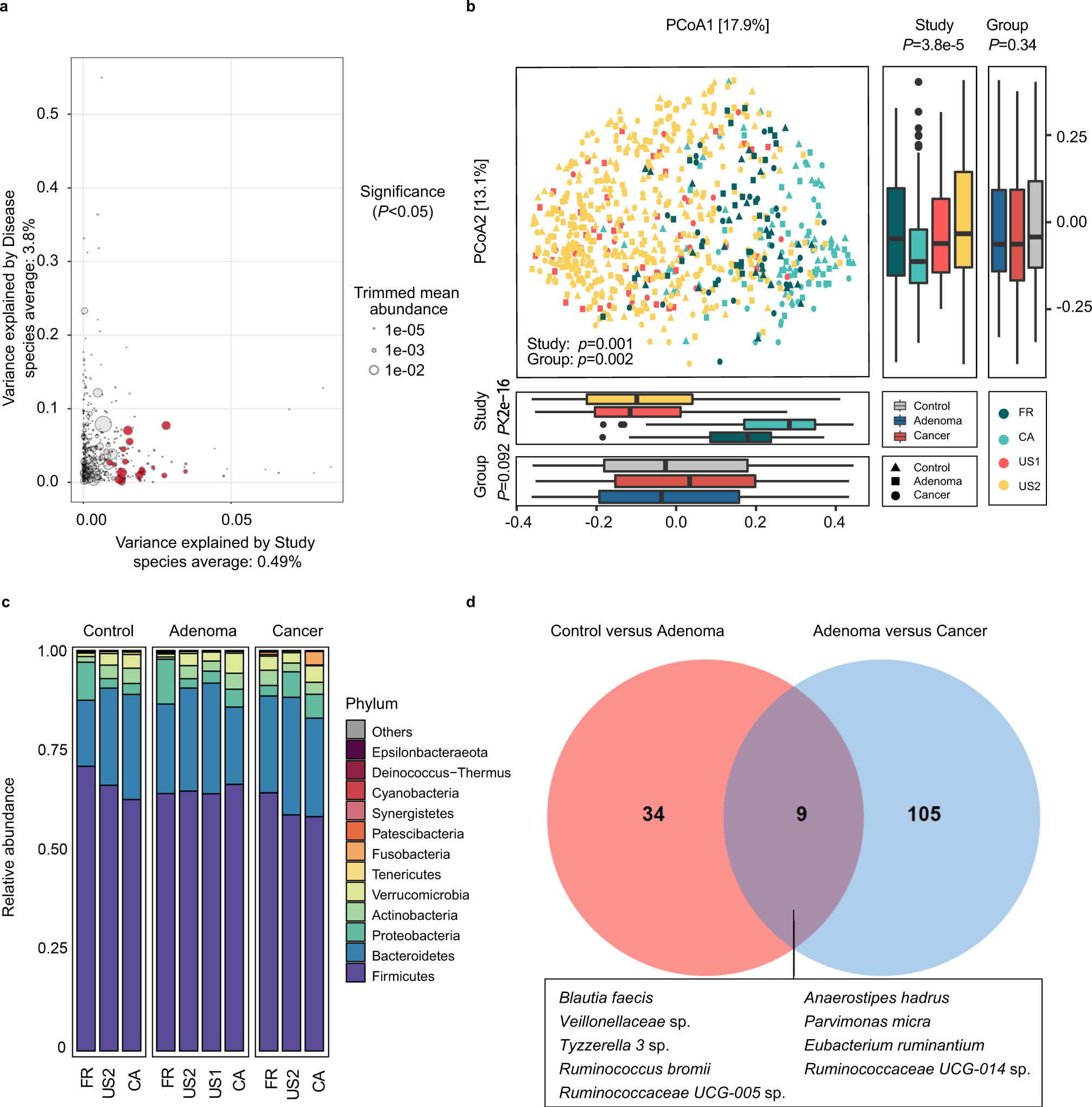
Identification of microbial markers across populations in early detection of colorectal cancer | Nature Communications
Regulation of Gene Expression in Autoimmune Disease Loci and the Genetic Basis of Proliferation in CD4+ Effector Memory T Cells | PLOS Genetics
RNA-Seq Signatures Normalized by mRNA Abundance Allow Absolute Deconvolution of Human Immune Cell Types
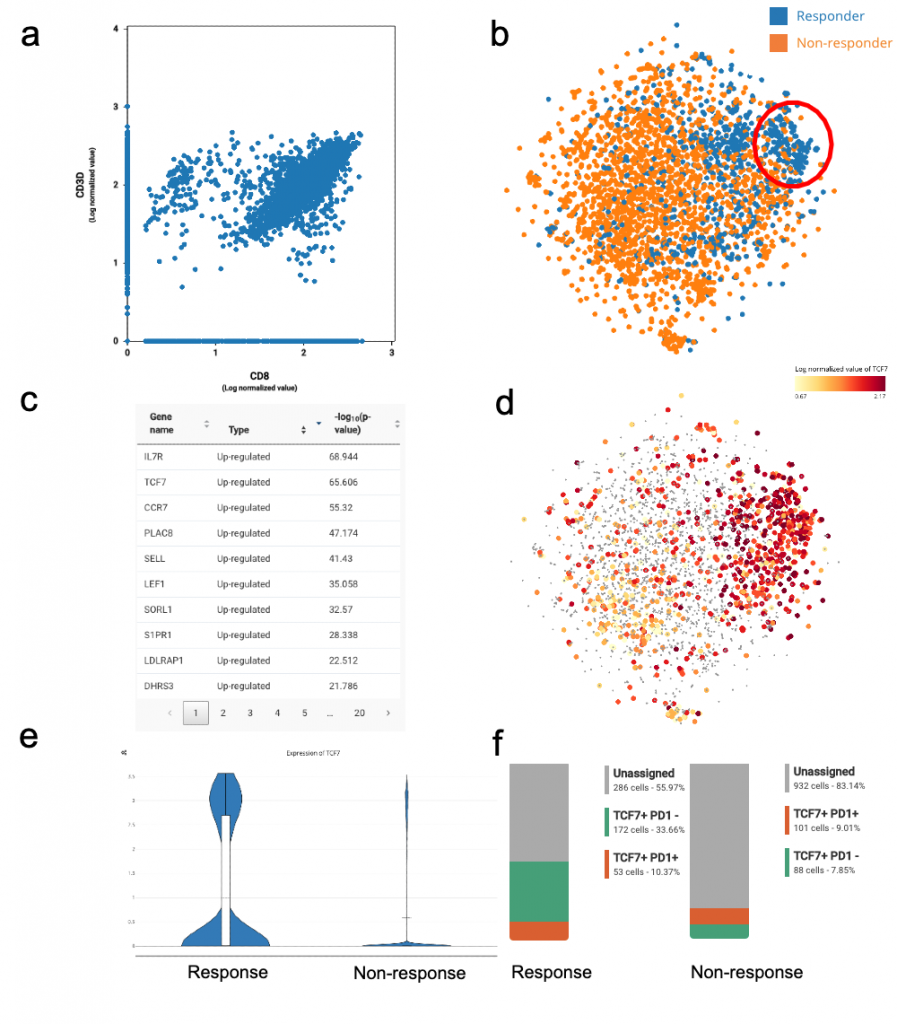
Single-cell analysis of CD8+ T cells in immune checkpoint blockade: some reproducible insights from BioTuring Database - BioTuring's Blog
![Genetic variability evaluation and cultivar identification of tetraploid annual ryegrass using SSR markers [PeerJ] Genetic variability evaluation and cultivar identification of tetraploid annual ryegrass using SSR markers [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2019/7742/1/fig-2-full.png)
Genetic variability evaluation and cultivar identification of tetraploid annual ryegrass using SSR markers [PeerJ]
Introduction to single-cell RNA-seq analysis - Differential expression and abundance between conditions

Single-cell analysis defines a pancreatic fibroblast lineage that supports anti-tumor immunity - ScienceDirect

A single-cell Arabidopsis root atlas reveals developmental trajectories in wild-type and cell identity mutants - ScienceDirect
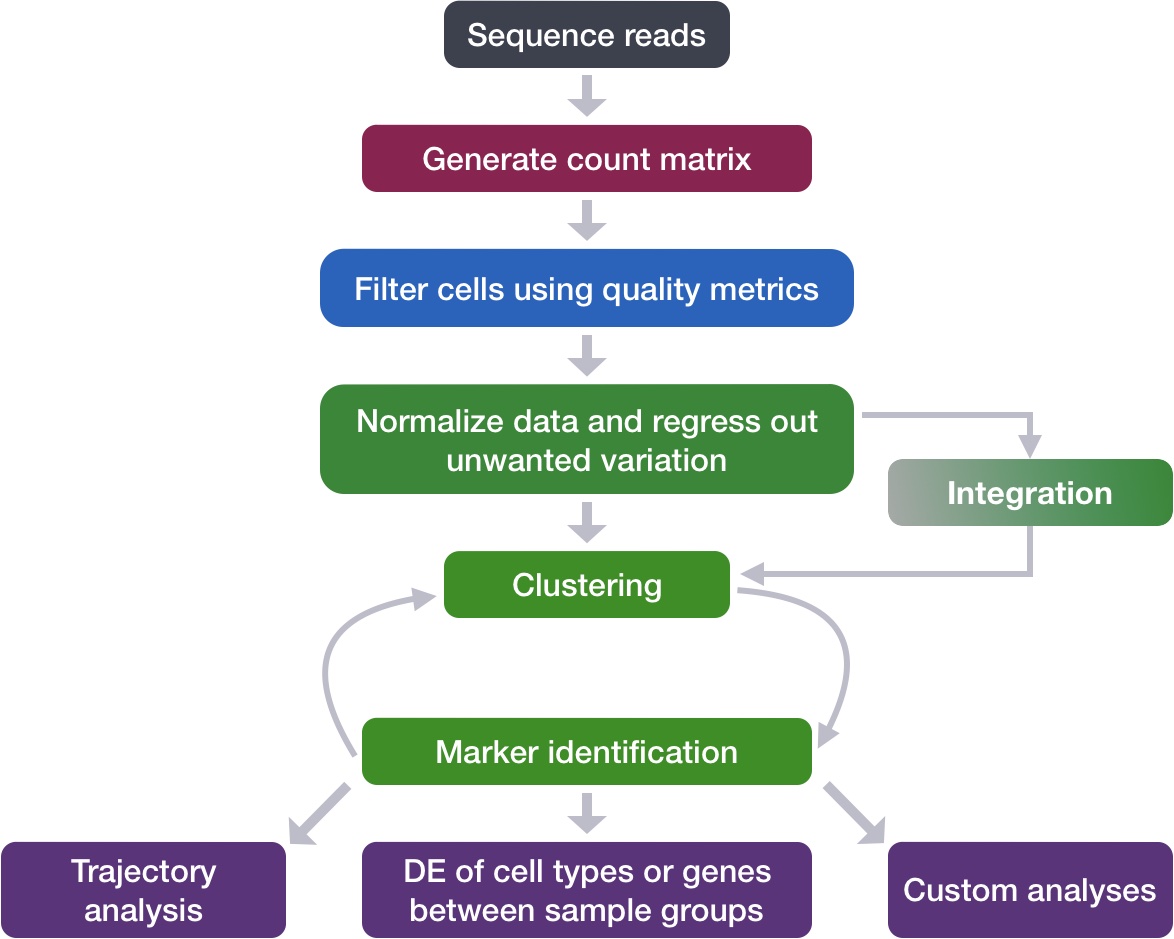
Single-cell RNA-seq: Normalization, identification of most variable genes | Introduction to single-cell RNA-seq



![Gene abundance (DNA) [gene copies] and gene expression (cDNA)... | Download Scientific Diagram Gene abundance (DNA) [gene copies] and gene expression (cDNA)... | Download Scientific Diagram](https://www.researchgate.net/publication/351854682/figure/fig3/AS:1027515707580418@1621990243168/Gene-abundance-DNA-gene-copies-and-gene-expression-cDNA-transcripts-of-the-marker.png)